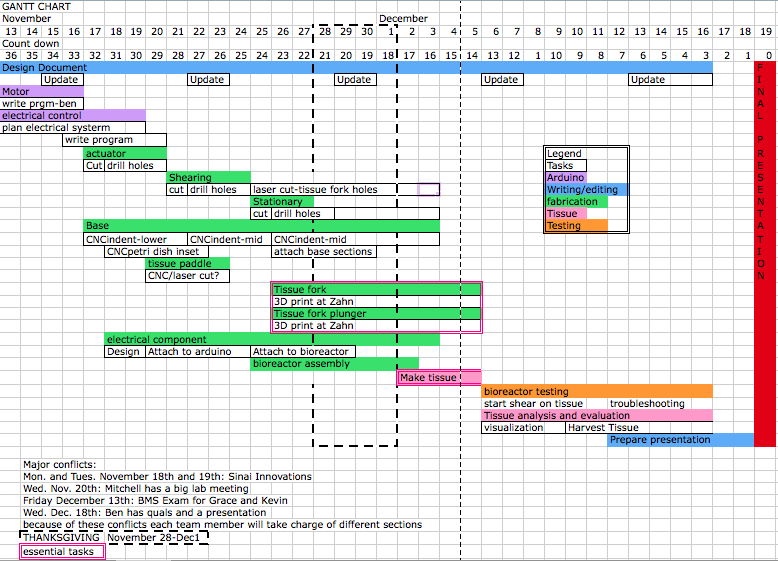
We’ve spent about a month at this point brainstorming ideas for our bioreactor and have settled on some initial goals and conclusions:
Our bioreactor will simulate compression on chondrocytes in cartilage. We picked compression because it seemed like a reasonable mechanical scenario to implement in the timespan we have, and it is relevant to clinical problems (e.g. repeated compression loading of intervertebral discs over time causes cellular damage, manifesting in herniations, osteoarthritis, etc.)
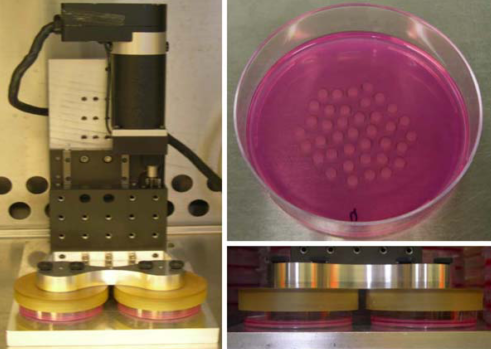
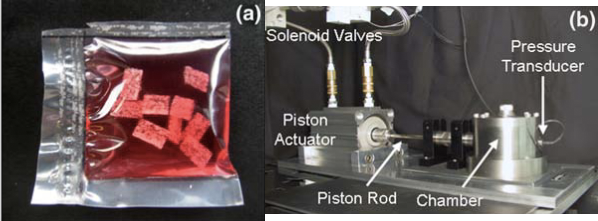
Currently, there are few high-throughput solutions for investigating drugs that can reverse the harmful effects of compression on articular cartilage. Bioreactors exist for applying hydrostatic pressure and compression force on 3D tissue cultures, but in general they deal with a small (1-10) number of conditions at once. See Mauck et al. 2007 for an example of a compression bioreactor, and Reza and Nicoll 2009 for an example of a hydrostatic pressure bioreactor. Relevant figures from each paper are below:
It is evident from these pictures that the experimental apparatus operates on dozens of replicates in the same environment, but there is no way to change the environment for individual replicates, nor to easily scale up the experiment to thousands of conditions. Very recent designs (see Hudson et al 2014) utilize 24-well cell culture plates but appear to require manual interaction for each experiment on each plate.
High throughput screening techniques have become more widely accessible to research laboratories as the cost of automating liquid/sample handling and imaging many samples at once has plummeted. The Integrated Screening Core at ISMMS is an example of one such facility that can automate the application of thousands of compounds to samples in 96- and 384-well plates, while reading fluorescence-based phenotypic assays. Most current screening experiments are based on phenotypes in 2D and liquid cell culture. Certain clinical scenarios however, like osteoarthritis or intervertebral disc herniation, are more realistically simulated in organs or 3D tissue culture, because the compression that occurs in vivo involves the 3D structure of the multicellular tissue.
Therefore, we wanted to make our bioreactor both able to compress 3D tissue cultures and integrate into a high throughput workflow, which we believe is a novel combination of capabilities. A major potential application of this work is the in vitro automated screening of up to thousands of compounds that reverse or mitigate injury phenotypes in chondrocytes, mesangial stem cells, and annulus fibrosus cells after repeated compression. Injury phenotypes that we could screen for in a relatively high-throughput manner using plate readers include apoptosis, as measured by a lanthanide-based flourescence assay for the activation of caspases. An example assay is described in Karvinen et al. 2004. Assay kits for caspases, including compatible microplate readers, are also commercially available. The discovery of compounds that can prevent compression injury in cartilage could have therapeutic potential for osteoarthritis, which affects 15% of the general population (Johnson and Hunter 2014), along with many other diseases of articular cartilage senescence and injury.